This notebook introduces the analysis of multi-temporal SAR backscatter image data stacks using the NISAR L2 GCOV data product. GCOV data are a geocoded data product that provides radiometric and terrain corrected backscatter measurements in at linear (power) scale.
This notebook demonstrates:
Accessing and loading of a GCOV times series from S3
Subsetting to an area of interest
Saving the subset time series as a local Zarr Store
Converting between power and sB scales
Generating a time series animation and saving it as a gif
Calculating a time series of mean backscatter.
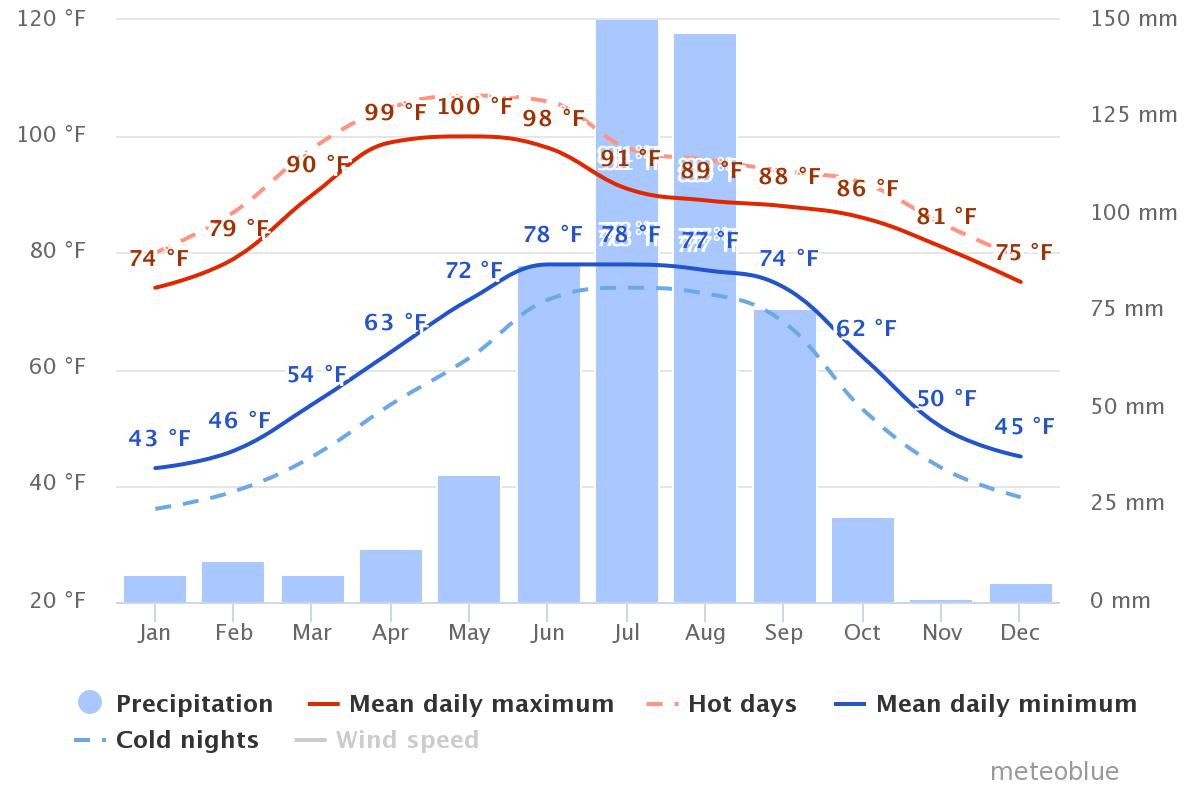
This notebook will be access a SAR data stack over Pokhara, Nepal for an introduction to time series processing. The data were acquired by NISAR’s L-band sensor and are available from the Alaska Satellite Facility.
Nepal is an interesting site for this analysis due to the significant seasonality of precipitation that is characteristic for this region. Nepal is said to have five seasons: spring, summer, monsoon, autumn and winter. Precipitation is low in the winter (November - March) and peaks dramatically in the summer, with top rain rates in July, August, and September (see figure to the right). As SAR is sensitive to changes in soil moisture, these weather patterns have a noticeable impact on the Radar Cross Section () time series information.
We will analyze the variation of values over time and will interpret them in the context of rainfall rates in the imaged area.
Overview¶
1. Prerequisites¶
| Prerequisite | Importance | Notes |
|---|---|---|
The isce3 software environment for this cookbook | Necessary | |
| How to search and access NISAR data in your AOI | Necessary | If you wish to process data in your AOI instead of the example data |
| Familiarity with xarray | Helpful | |
| Familiarity with matplotlib | Helpful |
Rough Notebook Time Estimate: 15 minutes (this is based on a 5 image time series and will change as more data become available)
2. Search Time Series Data¶
If you have run this notebook before and already saved a local copy of the time series, skip ahead to section 5.
2a. Search GCOV data and get a list of HTTPS product URLs with asf_search¶
import asf_search as asf
from datetime import datetime
import warnings
warnings.filterwarnings(
"ignore",
message="Parsing dates involving a day of month without a year specified",
)
session = asf.ASFSession()
start_date = datetime(2025, 11, 22)
end_date = datetime(2028, 1, 16)
area_of_interest = "POLYGON((83.8858 28.142,84.3554 28.142,84.3554 28.4825,83.8858 28.4825,83.8858 28.142))" # POINT or POLYGON as WKT (well-known-text)
pattern = r'^(?!.*QA_STATS).*' # used to remove static data files from results
opts=asf.ASFSearchOptions(**{
"maxResults": 250,
"intersectsWith": area_of_interest,
"start": start_date,
"end": end_date,
"processingLevel": [
"GCOV"
],
"dataset": [
"NISAR"
],
"productionConfiguration": [
"PR"
],
'session': session,
"frame": 16,
"relativeOrbit": 26,
"flightDirection": "ASCENDING",
})
response = asf.search(opts=opts)
hdf5_urls = response.find_urls(extension='.h5', pattern=pattern, directAccess=False)
print(f"Found {len(hdf5_urls)} GCOV products:")
hdf5_urlsFound 5 GCOV products:
['https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_026_A_016_4005_DHDH_A_20251123T234739_20251123T234814_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_026_A_016_4005_DHDH_A_20251123T234739_20251123T234814_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_007_026_A_016_4005_DHDH_A_20251205T234739_20251205T234814_X05009_N_F_J_001/NISAR_L2_PR_GCOV_007_026_A_016_4005_DHDH_A_20251205T234739_20251205T234814_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_008_026_A_016_4005_DHDH_A_20251217T234740_20251217T234815_X05009_N_F_J_001/NISAR_L2_PR_GCOV_008_026_A_016_4005_DHDH_A_20251217T234740_20251217T234815_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_009_026_A_016_4005_DHDH_A_20251229T234740_20251229T234815_X05009_N_F_J_001/NISAR_L2_PR_GCOV_009_026_A_016_4005_DHDH_A_20251229T234740_20251229T234815_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_010_026_A_016_4005_DHDH_A_20260110T234741_20260110T234816_X05009_N_F_J_001/NISAR_L2_PR_GCOV_010_026_A_016_4005_DHDH_A_20260110T234741_20260110T234816_X05009_N_F_J_001.h5']2b. Retain only the URL for the most recent version of each product in the search results¶
Data is occasionally re-released with an updated version. Versions are recorded as a Composite Release Identifier (CRID) in a product’s filename. We can use the CRID to retain only the most recent version of each product in the list of URLs.
import re
pattern = re.compile(r"(NISAR_L2_PR_GCOV(?:_[^_]+){9})_(X\d{5})")
latest_version_dict = {}
for url in hdf5_urls:
m = pattern.search(url)
if not m:
continue
product, crid = m.groups()
if product not in latest_version_dict or crid > latest_version_dict[product][0]:
latest_version_dict[product] = (crid, url)
hdf5_urls = [i[1] for i in latest_version_dict.values()]
print(f"Retained {len(hdf5_urls)} GCOV products:")
hdf5_urlsRetained 5 GCOV products:
['https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_026_A_016_4005_DHDH_A_20251123T234739_20251123T234814_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_026_A_016_4005_DHDH_A_20251123T234739_20251123T234814_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_007_026_A_016_4005_DHDH_A_20251205T234739_20251205T234814_X05009_N_F_J_001/NISAR_L2_PR_GCOV_007_026_A_016_4005_DHDH_A_20251205T234739_20251205T234814_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_008_026_A_016_4005_DHDH_A_20251217T234740_20251217T234815_X05009_N_F_J_001/NISAR_L2_PR_GCOV_008_026_A_016_4005_DHDH_A_20251217T234740_20251217T234815_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_009_026_A_016_4005_DHDH_A_20251229T234740_20251229T234815_X05009_N_F_J_001/NISAR_L2_PR_GCOV_009_026_A_016_4005_DHDH_A_20251229T234740_20251229T234815_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_010_026_A_016_4005_DHDH_A_20260110T234741_20260110T234816_X05009_N_F_J_001/NISAR_L2_PR_GCOV_010_026_A_016_4005_DHDH_A_20260110T234741_20260110T234816_X05009_N_F_J_001.h5']3b. Convert the HTTPS GCOV URLs into their S3 Bucket URLs¶
from urllib.parse import urlparse
bucket = "sds-n-cumulus-prod-nisar-products"
s3_urls = [f"s3://{bucket}/{'/'.join(urlparse(url).path.split('/')[2:])}" for url in hdf5_urls]
s3_urls['s3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_026_A_016_4005_DHDH_A_20251123T234739_20251123T234814_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_026_A_016_4005_DHDH_A_20251123T234739_20251123T234814_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_007_026_A_016_4005_DHDH_A_20251205T234739_20251205T234814_X05009_N_F_J_001/NISAR_L2_PR_GCOV_007_026_A_016_4005_DHDH_A_20251205T234739_20251205T234814_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_008_026_A_016_4005_DHDH_A_20251217T234740_20251217T234815_X05009_N_F_J_001/NISAR_L2_PR_GCOV_008_026_A_016_4005_DHDH_A_20251217T234740_20251217T234815_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_009_026_A_016_4005_DHDH_A_20251229T234740_20251229T234815_X05009_N_F_J_001/NISAR_L2_PR_GCOV_009_026_A_016_4005_DHDH_A_20251229T234740_20251229T234815_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_010_026_A_016_4005_DHDH_A_20260110T234741_20260110T234816_X05009_N_F_J_001/NISAR_L2_PR_GCOV_010_026_A_016_4005_DHDH_A_20260110T234741_20260110T234816_X05009_N_F_J_001.h5']3. Load the Data from S3 with xarray¶
3a. Get NISAR S3 Bucket credentials using your Earthdata Login Bearer Token¶
View or generate a Bearer Token in “Generate Token” tab of the Profile page in your Earthdata Login account: https://
from getpass import getpass
import json
import s3fs
import urllib
token = getpass("Enter your EDL Bearer Token")
prefix = "NISAR_L2_GCOV_BETA_V1"
event = {
"CredentialsEndpoint": "https://nisar.asf.earthdatacloud.nasa.gov/s3credentials",
"BearerToken": token,
"Bucket": bucket,
"Prefix": prefix,
}
# Get temporary download credentials
tea_url = event["CredentialsEndpoint"]
bearer_token = event["BearerToken"]
req = urllib.request.Request(
url=tea_url,
headers={"Authorization": f"Bearer {bearer_token}"}
)
with urllib.request.urlopen(req) as f:
creds = json.loads(f.read().decode())
fs = s3fs.S3FileSystem(
key=creds["accessKeyId"],
secret=creds["secretAccessKey"],
token=creds["sessionToken"],
)Enter your EDL Bearer Token ········
3b. Lazily load the Frequency A data for each GCOV product into an xarray.DataTree¶
Lazily load the frequencyA group of the GCOV data.
%%time
import xarray as xr
import rioxarray
# Explore the DataTree rendering above in Step 4 for a complete list of available groups
group_path = "/science/LSAR/GCOV/grids/frequencyA" # change this to any GCOV HDF5 group you wish
kwargs = {
"cache_type": "background",
"block_size": 16 * 1024 * 1024, # 16 MB
}
files = [fs.open(url, "rb", **kwargs) for url in s3_urls]
datatrees = [
xr.open_datatree(
f,
engine="h5netcdf",
decode_timedelta=False,
phony_dims="access",
chunks="auto",
group=group_path,
)
for f in files
]CPU times: user 1.11 s, sys: 388 ms, total: 1.5 s
Wall time: 6.03 s
3c. Create a list of datetimes for the time dimension of the time series¶
import re
from urllib.parse import urlparse
from datetime import datetime
from pathlib import PurePosixPath
NISAR_TS_RE = re.compile(r"_(\d{8}T\d{6})_")
def nisar_start_time_from_url(s3_url: str) -> datetime:
path = urlparse(s3_url).path
name = PurePosixPath(path).name
m = NISAR_TS_RE.search(name)
if not m:
raise ValueError(f"No NISAR timestamp found in: {s3_url}")
return datetime.strptime(m.group(1), "%Y%m%dT%H%M%S")
dts = [nisar_start_time_from_url(url) for url in s3_urls]
dts[datetime.datetime(2025, 11, 23, 23, 47, 39),
datetime.datetime(2025, 12, 5, 23, 47, 39),
datetime.datetime(2025, 12, 17, 23, 47, 40),
datetime.datetime(2025, 12, 29, 23, 47, 40),
datetime.datetime(2026, 1, 10, 23, 47, 41)]3d. Convert the Datatrees into DataArrays that include a time dimension¶
%%time
dataarrays = [
tree.ds.assign_coords(time=dt).expand_dims(time=1)
for dt, tree in zip(dts, datatrees)
]
dataarrays[0].timeCPU times: user 31.8 ms, sys: 238 μs, total: 32 ms
Wall time: 31.6 ms
3e. Extract the HHHH covariance layers and spatially subset them to an AOI¶
hhhh = [da.HHHH.rio.write_crs(f"EPSG:{da.isel(time=0).projection.item()}") for da in dataarrays]
bbox4326 = dict(minx=83.9633, miny=27.9427, maxx=84.3461, maxy=28.4555, crs="EPSG:4326")
hhhh_subset = [da.rio.clip_box(**bbox4326) for da in hhhh]ts = xr.concat(hhhh_subset, dim="time")
ts3f. Visually confirm the subset AOI’s location¶
Plot the subset bounds on a basemap to verify the expected location
import cartopy.crs as ccrs
import contextily as ctx
import matplotlib.pyplot as plt
import matplotlib.patches as patches
fig = plt.figure(figsize=(10,8))
ax = fig.add_subplot(1, 1, 1, projection=ccrs.epsg(3857))
minx, miny, maxx, maxy = hhhh[0].rio.transform_bounds("EPSG:3857", densify_pts=21)
ax.set_extent(
[minx, maxx, miny, maxy],
crs=ccrs.epsg(3857))
ctx.add_basemap(ax, source=ctx.providers.Esri.WorldImagery)
ctx.add_basemap(ax, source=ctx.providers.OpenStreetMap.Mapnik, alpha=0.5)
sub_minx, sub_miny, sub_maxx, sub_maxy = hhhh_subset[0].rio.transform_bounds("EPSG:3857", densify_pts=21)
rect = patches.Rectangle(
(sub_minx, sub_miny),
sub_maxx - sub_minx,
sub_maxy - sub_miny,
linewidth=2,
edgecolor="red",
facecolor="none",
label="Subset AOI",
transform=ccrs.epsg(3857)
)
ax.add_patch(rect)
ax.legend(handles=[rect])
plt.show()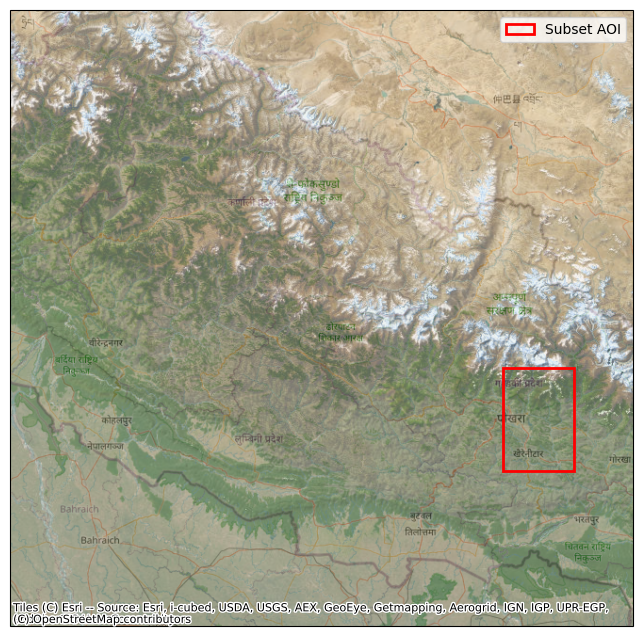
4. (Optional) Save the time series as a local Zarr Store¶
Up to this point, you have lazily loaded the time series data into xarray data structures. These hold only pointers to data stored remotely in an S3 Bucket. Accessing the values in a large dataset from S3 in Earthdata can be slow. If you are not careful in how you work with the them, you may access the same data multiple times, slowing your work even more. Additionally, Earthdata S3 Bucket credentials expire after one hour, so if you don’t complete your work within that time, you may have to repeat many slow steps in your analysis.
To mitigate these pain points, you can save your subset time series to a Zarr Store on your local volume.
Local data access is much faster than remote S3 access.
It is still wise to avoid recomputing values via careful use of xarray.
If you set down your work for the day or restart your notebook kernel or JupyterHub server, you can quickly reload your locally stored data.
You are no longer time-constrained by short-lived Earthdata S3 Bucket credentials.
4a. Create directories to hold the time series data and output plots¶
from pathlib import Path
time_series_example_dir = Path.home() / "NISAR_GCOV_time_series_example"
data_dir = time_series_example_dir / "data"
data_dir.mkdir(parents=True, exist_ok=True)4b. Save the time series as a Zarr Store¶
Until now, we have not accessed any values in our xarray data structures; we have manipulated pointers to the data. To save the data to a Zarr Store, all values in the subset must be accessed, which is slow over the network.
%%time
ts = ts.chunk({"time": 1, "yCoordinates": 2048, "xCoordinates": 2048})
zarr_ts_path = data_dir / "pokhara_timeseries.zarr"
ts.to_zarr(data_dir / "pokhara_timeseries.zarr", mode="w", consolidated=False)CPU times: user 12.2 s, sys: 3.65 s, total: 15.9 s
Wall time: 47.4 s
<xarray.backends.zarr.ZarrStore at 0x7f568c4949a0>4c. Close the open files to the data in S3¶
for f in files:
f.close()5. (Optional) Load the time series from the local Zarr Store¶
If you have opted to run Step 4 of this notebook and saved a local Zarr Store, you can start the notebook from from this step when re-running it.
5a. Load the Zarr Store¶
%%time
from pathlib import Path
import xarray as xr
time_series_example_dir = Path.home() / "NISAR_GCOV_time_series_example"
data_dir = time_series_example_dir / "data"
plot_dir = time_series_example_dir / "plots"
zarr_ts_path = data_dir / "pokhara_timeseries.zarr"
ts = xr.open_zarr(zarr_ts_path, consolidated=False).HHHH
tsCPU times: user 2.82 ms, sys: 10.9 ms, total: 13.8 ms
Wall time: 12.1 ms
6. Data Conversion between dB and Power Scales¶
Focused SAR image data may come in uncalibrated digital numbers (DN) and need to be calibrated to correspond to proper radar cross section information. However, NISAR data is already radiometrically calibrated and has a scaling factor of 1.0, so we do not need to worry about this.
The data at hand are radiometrically terrain corrected images, which are expressed as terrain flattened backscatter coefficients. For forest and land cover monitoring applications is the preferred metric.
We often visualize backscatter in log scale (dB), as plotting in a linear (Power) scale may obscure subtleties. However, we must compute statistics in linear scale, so you may have need to convert back and forth between Power and dB.
NISAR GCOV data are provided in Power scale.
6a. Convert the HHHH data from power to dB¶
To convert from power to dB, apply:
%%time
import numpy as np
ts_db = db = 10 * np.log10(ts.where(ts > 0)) # avoid log10 on values <= 0 to avoid division by zero warnings
# load ts_db values into memory once now to avoid multiple loads in subsequent code cells
ts_db.load()
ts_dbCPU times: user 1.96 s, sys: 307 ms, total: 2.27 s
Wall time: 690 ms
6b. Plot the histogram of the dB data and confirm a normal distribution¶
import matplotlib.pyplot as plt
ax = ts_db.values.ravel()
ax = ax[(ax < 20) & (ax > -40)]
plt.hist(ax, bins=50)
plt.show()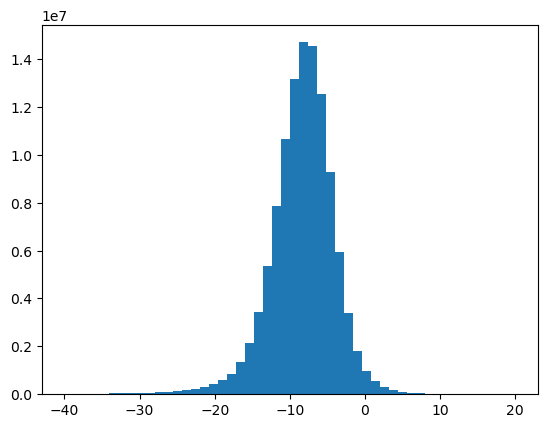
6c. Convert from dB back to power¶
While dB-scaled images are often visually pleasing, they are not a good basis for mathematical operations. For instance, when computing the mean backscatter, we cannot simply average data in log scale.
Please note that the correct scale in which operations need to be performed is the power scale. This is critical, e.g. when speckle filters are applied, when spatial operations like block averaging are performed, or other statistics across time series are analyzed.
To convert from dB to power, apply:
ts_pwr = np.power(10.0, ts_db/10.0)
ts_pwrPlot the histogram of the Power-scale data¶
ax = ts_pwr.values.ravel()
ax = ax[ax < 2]
plt.hist(ax, bins=500)
plt.show()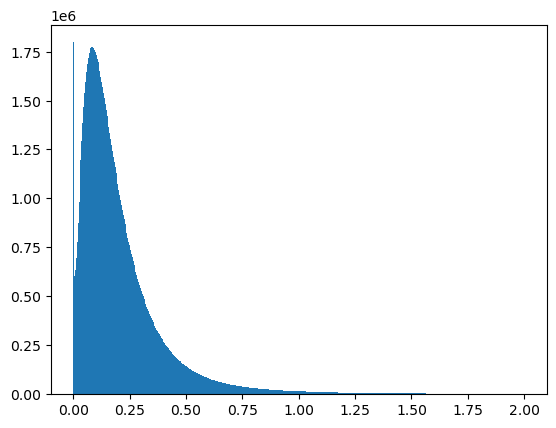
7. SAR Time Series Animation¶
This section introduces you to the handling and analysis of SAR time series stacks. A focus will be put on time series visualizations, which allow us to inspect time series in more depth.
Animations will display interactively and be saved as gifs.
7a. Create a Time Series Animation from the dB Subset¶
%%capture
from matplotlib.animation import FuncAnimation
from matplotlib.animation import FFMpegWriter, PillowWriter
vmin = ts_db.quantile(0.05, skipna=True).values.item()
vmax = ts_db.quantile(0.95, skipna=True).values.item()
fig, ax = plt.subplots(figsize=(6, 6), )
frame_0 = ts_db.isel(time=0).values
im = ax.imshow(frame_0, vmin=vmin, vmax=vmax, origin="upper", cmap="grey")
cb = fig.colorbar(im, ax=ax, label="γ⁰ (dB)")
ax.set_title(np.datetime_as_string(ts_db.time.values[0], unit='s'))
ax.set_axis_off()
def animate(i):
frame = ts_db.isel(time=i).values
im.set_data(frame)
ax.set_title(np.datetime_as_string(ts_db.time.values[i], unit='s'))
return (im,)
ani = FuncAnimation(fig, animate, frames=ts_db.sizes["time"], interval=300, blit=False)
plt.show()7b. Create a javascript animation of the time-series running inline in the notebook:¶
from IPython.display import HTML
HTML(ani.to_jshtml())7c. Save the animation (animation.gif):¶
from pathlib import Path
time_series_example_dir = Path.home() / "NISAR_GCOV_time_series_example"
plot_dir = time_series_example_dir / "plots"
plot_dir.mkdir(parents=True, exist_ok=True)
ani.save(plot_dir / "timeseries_animation.gif", writer=PillowWriter(fps=3))8. Calculate the Time Series of Means¶
Calculate the Time Series of Means Across the HHHH
To create the time series of means:
Compute subset means for each time-step using data in power scale () .
Convert the resulting mean values into dB scale for visualization.
8a. Identify the axis on which to take the means¶
Note that the Dataset dimensions are (‘time’, ‘yCoordinates’, ‘xCoordinates’)
ts_pwr.dims('time', 'yCoordinates', 'xCoordinates')8b. Compute the means in power scale:¶
Since we want to take the means across the spatial dimensions (dims 1 and 2) on each day, we should pass axis=(1, 2) to np.mean.
This will compute the backscatter means spatially, while leaving the time dimension in place.
means_pwr = np.mean(ts_pwr, axis=(1, 2))
means_pwr8c. Convert the resulting mean value time-series to dB scale for visualization¶
means_db = 10 * np.log10(means_pwr.where(means_pwr > 0))
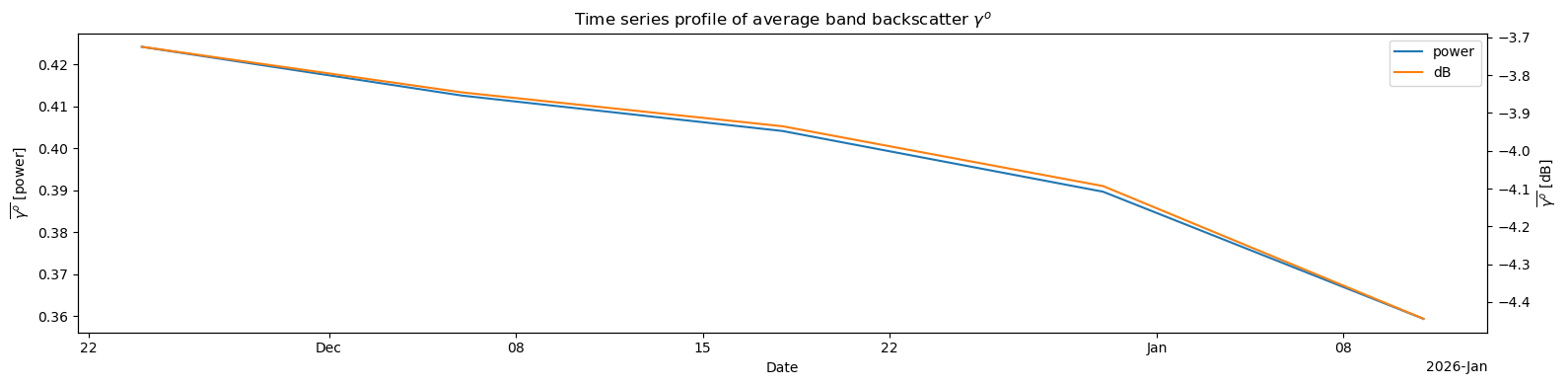
means_db8d. Plot the time series of means¶
Plot both the Power and dB time series of means
import matplotlib.dates as mdates
fig, ax1 = plt.subplots(figsize=(16, 4))
# left y-axis: power
means_pwr.plot.line(ax=ax1, x="time", label="power")
ax1.set_xlabel("Date")
ax1.set_ylabel(r"$\overline{\gamma^o}$ [power]")
# right y-axis: dB
ax2 = ax1.twinx()
means_db.plot.line(ax=ax2, x='time', color='tab:orange', label='dB')
ax2.set_ylabel(r'$\overline{\gamma^o}$ [dB]')
ax2.set_xlim(ax1.get_xlim())
h1, l1 = ax1.get_legend_handles_labels()
h2, l2 = ax2.get_legend_handles_labels()
ax1.legend(h1 + h2, l1 + l2, loc='upper right')
ax1.set_title(r'Time series profile of average band backscatter $\gamma^o$')
ax2.set_title("")
plt.tight_layout()
plt.show()
8e. Create Two-Panel Figure, Animating Both the Imagery and the Daily Global Means ¶
%%capture
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.animation as an
import matplotlib.dates as mdates
vmin = ts_db.quantile(0.05, skipna=True).values.item()
vmax = ts_db.quantile(0.95, skipna=True).values.item()
t_num = mdates.date2num(ts_db.time.values)
means = np.asarray(means_db.compute().values).reshape(-1)
fig = plt.figure(figsize=(12,4))
ax1 = fig.add_subplot(121)
ax2 = fig.add_subplot(122)
# Image axis
frame_0 = ts_db.isel(time=0).values
im = ax1.imshow(frame_0, vmin=vmin, vmax=vmax, origin="upper", cmap="grey")
ax1.set_title(np.datetime_as_string(ts_db.time.values[0], unit='D'))
ax1.set_axis_off()
# Line axis
(l,) = ax2.plot([], [], lw=3)
ax2.xaxis.set_major_formatter(mdates.DateFormatter('%Y-%m-%d'))
ax2.set_xlim(t_num.min(), t_num.max())
ax2.set_ylim(np.nanmin(means)-0.5, np.nanmax(means)+0.5)
ax2.set_ylabel(r"Backscatter Means $\mu_{\gamma^0_{dB}}$")
ax2.set_title(str(ts_db.time.values[0].astype("datetime64[D]")))
def init():
l.set_data([], [])
return im, l
def animate(i):
ts = str(ts_db.time.values[i].astype("datetime64[D]"))
ax1.set_title(ts)
ax2.set_title(ts)
im.set_data(ts_db.isel(time=i).values)
l.set_data(t_num[:i+1], means[:i+1])
return im, l
ani = an.FuncAnimation(fig, animate, frames=len(ts_db.time), init_func=init, interval=400, blit=False)
plt.tight_layout()
plt.show()8f. Create a javascript animation of the time-series running inline in the notebook:¶
HTML(ani.to_jshtml())Save the animated time-series and histogram (animation_histogram.gif):¶
from pathlib import Path
time_series_example_dir = Path.home() / "NISAR_GCOV_time_series_example"
plot_dir = time_series_example_dir / "plots"
plot_dir.mkdir(parents=True, exist_ok=True)
ani.save(plot_dir / 'animation_histogram.gif', writer='pillow', fps=2)9. Summary¶
Having completed this workflow, you have:
Searched and downloaded a NISAR GCOV time series
Subset the data to a specific polarization and AOI
Converted the data between linear and logarithmic scales
Generated a backscatter time series animation and output it as a gif
Calculated the time series of means, animated it, and output it as a gif
10. Resources and references¶
Authors:
Alex Lewandowski; Alaska Satellite Facility
Adapted for NISAR data from earlier work by:
Franz J Meyer; University of Alaska Fairbanks
Josef Kellndorfer, Earth Big Data, LLC