NISAR Pixel Conversion¶
This notebook converts sample ground truth data coordinates (latitude and longitude) into corresponding NISAR raster pixel indices. Identifying the correct pixel location for each site allows direct extraction of NISAR data values at field observation points.
The workflow includes transforming geographic coordinates into the NISAR product’s coordinate system, locating the nearest raster pixel, and storing the associated row and column indices for analysis. The resulting site locations and pixel footprints are also visualized on an interactive Folium map to confirm alignment between ground sites and the NISAR raster grid.
Ground Truth Site¶
About Lake Afrera¶
Location and Setting
Lake Afrera (also known as Lake Giulietti) is located in northeastern Ethiopia within the Danakil Depression, one of the lowest and hottest places on Earth. The lake sits in the Afar Region, near the border with Eritrea, at an elevation of approximately −100 meters below sea level.
The Danakil Depression lies within the Afar Triple Junction, where the African, Arabian, and Somali tectonic plates are actively pulling apart. This rifting environment creates intense volcanic and geothermal activity throughout the region.
Why Lake Afrera Is Hypersaline?
Lake Afrera is a hypersaline lake, meaning its salinity is significantly higher than that of normal seawater. Its high salinity results from several factors:
Closed basin hydrology: The lake has no outlet to the ocean, so dissolved salts accumulate over time.
Extreme evaporation: The Danakil Depression experiences very high temperatures and low rainfall, causing water to evaporate rapidly.
Hydrothermal inputs: Nearby geothermal systems and volcanic activity contribute mineral-rich fluids.
As water evaporates, salts such as halite (sodium chloride) precipitate and form crusts and salt flats around the lake margins. These processes create strong contrasts in surface roughness and reflectivity, which are visible in radar imagery.
Surface Features and Radar Relevance
Lake Afrera and its surrounding landscape display strong contrasts in surface texture that are particularly visible in radar data. The region includes open water (typically dark in radar imagery), bright salt crusts and evaporite deposits, mud flats that vary seasonally, and geometric salt extraction ponds.
The lake is also bordered by volcanic cones, basaltic lava flows, and geothermal areas associated with the Afar rift system. This mix of water, salt, sediment, and volcanic rock creates a highly heterogeneous surface within a relatively small area.
These sharp contrasts in roughness, moisture, and dielectric properties make Lake Afrera an excellent natural laboratory for studying radar backscatter variability and polarization-dependent scattering behavior.
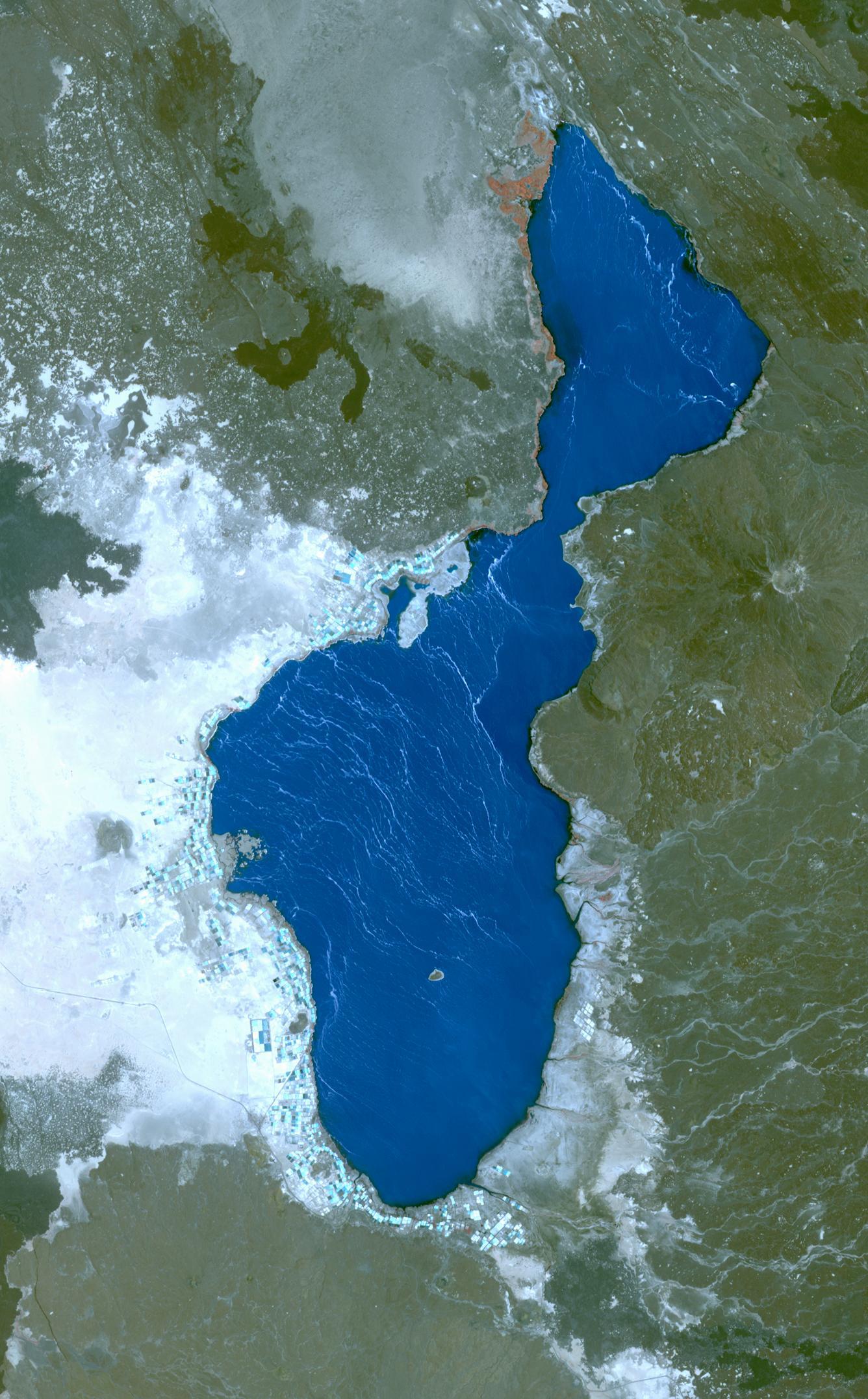
Optical satellite image of Lake Afrera and surrounding geolgical features: https://
Notebook Overview¶
1. Prerequisites¶
| Prerequisite | Importance | Notes |
|---|---|---|
| The software environment for this cookbook must be installed | Necessary |
Rough Notebook Time Estimate: 10 minutes
import os
import asf_search as asf
from datetime import datetime
from getpass import getpass
from pathlib import Path
#1: Search for and download NISAR GCOV HDF5 products from ASF
#authenticate with Earthdata Login
#create an ASF session object (stores your login + settings for requests)
session = asf.ASFSession()
#prompt for username/password
session.auth_with_creds(input('EDL Username'), getpass('EDL Password'))
#2: Define the search window and area of interest
#insert start and end dates. Note that dates and times are inclusive.
start_date = datetime(2025, 12, 4)
end_date = datetime(2025, 12, 5)
# POINT or POLYGON as WKT (well-known-text) using lat/lon pairs
area_of_interest = "POLYGON((40.9131 12.3904,41.8891 12.3904,41.8891 13.2454,40.9131 13.2454,40.9131 12.3904))"
#3: Build search options
#these filters narrow results to NISAR GCOV products over the AOI and time window
opts = asf.ASFSearchOptions(
**{
"maxResults": 250, # cap the number of returned results
"intersectsWith": area_of_interest, # spatial filter (WKT geometry)
"start": start_date, # temporal filter (start)
"end": end_date, # temporal filter (end)
"processingLevel": ["GCOV"], # product type / processing level
"dataset": ["NISAR"], # mission/dataset name
"productionConfiguration": ["PR"], # production config (e.g., provisional)
"session": session, # use authenticated session from above
}
)
#4: Run the search for HDF5 files
#execute search
response = asf.search(opts=opts)
#exclude QA_STATE files
pattern = r'^(?!.*QA_STATS).*'
#extract files that end with .h5
hdf5_files = response.find_urls(extension='.h5', pattern=pattern, directAccess=False)
hdf5_filesEDL Username aflewandowski
EDL Password ········
['https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05007_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05007_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001.h5']2b. Retain only the URL for the most recent version of each product in the search results¶
Data is occasionally re-released with an updated version. Versions are recorded as a Composite Release Identifier (CRID) in a product’s filename. We can use the CRID to retain only the most recent version of each product in the list of URLs.
import re
pattern = re.compile(r"(NISAR_L2_PR_GCOV(?:_[^_]+){9})_(X\d{5})")
latest_version_dict = {}
for url in hdf5_files:
m = pattern.search(url)
if not m:
continue
product, crid = m.groups()
if product not in latest_version_dict or crid > latest_version_dict[product][0]:
latest_version_dict[product] = (crid, url)
hdf5_files = [i[1] for i in latest_version_dict.values()]
print(f"Retained {len(hdf5_files)} GCOV products:")
hdf5_filesRetained 1 GCOV products:
['https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001.h5']2c. Download the data¶
data_dir = Path.home() / "GCOV_data_pixel_conversion_example"
data_dir.mkdir(exist_ok=True)
print(f'data_dir: {data_dir}')
#download files
asf.download_urls(hdf5_files, data_dir, session=session, processes=4)data_dir: /home/jovyan/GCOV_data_pixel_conversion_example
3. Locate Relevant GCOV data¶
Navigate to the grids group in the HDF5 file to select frequency and polarization.
import h5py
#path to the NISAR GCOV HDF5 file
h5_path = list(data_dir.glob("NISAR_L2_*_GCOV*.h5"))[0]
#GCOV data are stored under the "grids" groups
#select desired frequency: "frequencyA" or "frequencyB"
freq = "frequencyA"
grid_grp = f"/science/LSAR/GCOV/grids/{freq}"
#for this example, we will be using the HHHHH layer
#for this GCOV file, different layers are available and listed in the output below.
layer = "HHHH"
layer_path = f"{grid_grp}/{layer}"
#print paths
print("File:", h5_path)
print("Grid group:", grid_grp)
print("Current raster layer:", layer_path)
#examine available GCOV datasets within the file:
with h5py.File(h5_path, "r") as f:
print("\nAvailable GCOV datasets:")
for k in f[grid_grp].keys():
print(" ", k)File: /home/jovyan/GCOV_data_pixel_conversion_example/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001.h5
Grid group: /science/LSAR/GCOV/grids/frequencyA
Current raster layer: /science/LSAR/GCOV/grids/frequencyA/HHHH
Available GCOV datasets:
HHHH
HVHV
listOfCovarianceTerms
listOfPolarizations
mask
numberOfLooks
numberOfSubSwaths
projection
rtcGammaToSigmaFactor
xCoordinateSpacing
xCoordinates
yCoordinateSpacing
yCoordinates
4. Identify the coordinate system used by GCOV¶
Extract coordinate system, raster size, and pixel size.
#open the GCOV file (in read mode) to read coordinate and spatial information
with h5py.File(h5_path, "r") as f:
#in a GCOV file, the "projection" dataset stores coordinate systems
#the output value will be an EPSG code
epsg = int(f[f"{grid_grp}/projection"][()])
#xCoordinates and yCoordinates are arrays that store the coordinates of each pixel center
#these coordinates will later be used to center pixel footprints
x = f[f"{grid_grp}/xCoordinates"][:]
y = f[f"{grid_grp}/yCoordinates"][:]
#CoordinateSpacing values store the pixel size in meters
#this will later be used to map the pixel footprint
dx = float(f[f"{grid_grp}/xCoordinateSpacing"][()])
dy = float(f[f"{grid_grp}/yCoordinateSpacing"][()])
#print information
print("GCOV coordinate system (EPSG):", epsg)
print("Raster size:", len(y), "rows x", len(x), "columns")
print("Pixel size:", dx, "m (x) ×", dy, "m (y)")GCOV coordinate system (EPSG): 32637
Raster size: 16704 rows x 17064 columns
Pixel size: 20.0 m (x) × -20.0 m (y)
5. Convert ground truth site locations to GCOV pixel coordinates¶
5a. Define ground truth site coordinates¶
#define sites
#these coordinates are in EPSG 4326
sites = {
"Salt_Ponds": (13.25889, 40.85111),
"Hypersaline_Lake": (13.26028, 40.89972),
"Afdera_Volcano": (13.08833, 40.85250),
"Sand_Plains": (13.27944, 40.80694),
"Afdera_Lava_Field": (13.09222, 40.72583),
"Borawli_Caldera": (13.30389, 40.98972),
"Borawli_Lava_Field": (13.21833, 40.98056),
}5b. Convert Lake Afrera site coordinates to GCOV pixel coordinates.¶
from pyproj import Transformer
import numpy as np
#use transformer to convert ground truth data (EPSG 4326) coordinate system to GCOV coordinate system (EPSG 32637)
transformer = Transformer.from_crs(
"EPSG:4326",
f"EPSG:{epsg}",
always_xy=True
)
site_pixels = {}
for name, (lat, lon) in sites.items():
#convert the site location into GCOV map coordinates
x_site, y_site = transformer.transform(lon, lat)
#find the nearest GCOV pixel by comparing with the grid coordinates
col = int(np.argmin(np.abs(x - x_site)))
row = int(np.argmin(np.abs(y - y_site)))
#store the GCOV pixel index for this site
site_pixels[name] = (row, col)
#GCOV pixel indices corresponding to each site
site_pixels{'Salt_Ponds': (6090, 4823),
'Hypersaline_Lake': (6080, 5087),
'Afdera_Volcano': (7034, 4838),
'Sand_Plains': (5978, 4583),
'Afdera_Lava_Field': (7017, 4151),
'Borawli_Caldera': (5835, 5573),
'Borawli_Lava_Field': (6309, 5527)}6. Create Folium map¶
This map will contain:
HHHH overlay + HVHV overlay
ground truth site markers (blue)
GCOV pixel footprints (yellow)
Additional tools: measure tool, mouse coordinates, layer toggle
This code uses the following variables from previous cells:
h5_path, grid_grp, epsg, x, y, dx, dy, sites, site_pixels
import folium
from folium.raster_layers import ImageOverlay
from folium.plugins import MeasureControl, MousePosition
import matplotlib.pyplot as plt
import branca.colormap as cm
################## Display / quicklook settings
#downsample factor for faster visualization
#the "step" variable tells Python to read every Nth pixel in both row and column directions
step = 60
#overlay transparency (0 = invisible, 1 = opaque)
overlay_opacity = 0.55
################# Convert coordinates into lat/lon for Folium use
# Folium expects WGS84 lat/lon (EPSG:4326).
to_ll = Transformer.from_crs(f"EPSG:{epsg}", "EPSG:4326", always_xy=True)
################ Read downsampled arrays (HHHH + HVHV) for quicklooks
#note: this does NOT change the original data in the HDF5 file
with h5py.File(h5_path, "r") as f:
#read downsampled polarization arrays for display
hh_q = np.array(f[f"{grid_grp}/HHHH"][::step, ::step], dtype=float)
hv_q = np.array(f[f"{grid_grp}/HVHV"][::step, ::step], dtype=float)
#read downsampled coordinate arrays (meters)
#these are used to place the overlay in the correct geographic location
xq = f[f"{grid_grp}/xCoordinates"][::step]
yq = f[f"{grid_grp}/yCoordinates"][::step]
################# Stretch display settings to allow easier visualization
#this affects visualization only — the original GCOV data are unchanged.
#using a single shared vmin/vmax ensures that both polarizations use the same
#brightness scale, preventing misleading visual comparisons between HHHH and HVHV
stack = np.concatenate([hh_q[np.isfinite(hh_q)], hv_q[np.isfinite(hv_q)]])
vmin = np.nanpercentile(stack, 2)
vmax = np.nanpercentile(stack, 98)
print("Stretched display range (2–98% percentile):")
print(f"vmin = {vmin}")
print(f"vmax = {vmax}\n")
################# Convert GCOV rasters to RGBA image
#folium cannot overlay the GCOV rasters in their current format, so they must
#be converted to RGBA images for visualization
def arr_to_rgba(arr, vmin, vmax):
#mask missing values
finite = np.isfinite(arr)
#clip to display range
arr_clip = np.clip(arr, vmin, vmax)
#normalize to 0–1 for colormap
norm = (arr_clip - vmin) / (vmax - vmin + 1e-12)
norm = np.nan_to_num(norm, nan=0.0)
#map to RGBA using grayscale
rgba = plt.cm.gray(norm)
#set alpha channel to transparent for NaNs
rgba[..., 3] = finite.astype(float) * 0.85
return rgba
#convert both polarizations to RGBA using the same stretch
rgba_hh = arr_to_rgba(hh_q, vmin, vmax)
rgba_hv = arr_to_rgba(hv_q, vmin, vmax)
################# Compute overlay bounds (lat/lon) from GCOV x/y (meters)
#ImageOverlay requires SW/NE bounds in lat/lon
xmin, xmax = float(xq.min()), float(xq.max())
ymin, ymax = float(yq.min()), float(yq.max())
lon_sw, lat_sw = to_ll.transform(xmin, ymin)
lon_ne, lat_ne = to_ll.transform(xmax, ymax)
bounds = [[lat_sw, lon_sw], [lat_ne, lon_ne]]
################# Center Folium map over ground truth sites
center_lat = sum(lat for lat, lon in sites.values()) / len(sites)
center_lon = sum(lon for lat, lon in sites.values()) / len(sites)
m = folium.Map(
location=[center_lat, center_lon],
zoom_start=8,
tiles="Esri.WorldImagery"
)
################# Add toggleable overlays
ImageOverlay(
image=rgba_hh,
bounds=bounds,
opacity=overlay_opacity,
name="HHHH",
interactive=True,
zindex=2
).add_to(m)
ImageOverlay(
image=rgba_hv,
bounds=bounds,
opacity=overlay_opacity,
name="HVHV",
interactive=True,
zindex=2
).add_to(m)
################# Add ground truth sites and GCOV pixel footprints to map
sites_fg = folium.FeatureGroup(name="Ground Truth Sites", show=True)
pix_fg = folium.FeatureGroup(name="GCOV Pixel Footprints", show=True)
for name, (lat, lon) in sites.items():
#groundn truth site marker (blue)
folium.Marker(
location=[lat, lon],
popup=f"{name}",
icon=folium.Icon(color="blue")
).add_to(sites_fg)
#pixel footprint (yellow)
row, col = site_pixels[name]
#pixel center coordinates
x_center = float(x[col])
y_center = float(y[row])
#pixel corners (in meters)
corners_xy = [
(x_center - dx/2, y_center - dy/2),
(x_center + dx/2, y_center - dy/2),
(x_center + dx/2, y_center + dy/2),
(x_center - dx/2, y_center + dy/2),
(x_center - dx/2, y_center - dy/2),
]
#convert corners to lat/lon for folium
corners_latlon = []
for xc, yc in corners_xy:
lonp, latp = to_ll.transform(xc, yc)
corners_latlon.append([latp, lonp])
folium.Polygon(
locations=corners_latlon,
color="yellow",
weight=2,
fill=False,
popup=f"{name}"
).add_to(pix_fg)
sites_fg.add_to(m)
pix_fg.add_to(m)
################# Add Legend
colormap = cm.linear.Greys_09.scale(round(vmin, 5), round(vmax, 5))
colormap.caption = "GCOV backscatter (shared stretch)"
# colormap.add_to(m)
################# Add interactive tools
MeasureControl(position="topleft").add_to(m) #measure distance/area
MousePosition(position="bottomleft").add_to(m) #display lat/lon for cursor
m.add_child(folium.LatLngPopup()) #click map to get lat/lon popup
#keep control panel open
folium.LayerControl(collapsed=False).add_to(m)
display(colormap)
display(m)Stretched display range (2–98% percentile):
vmin = 0.0009728094935417175
vmax = 0.31600227355957017
Summary¶
You have now opened and navigated an HDF5 file, and converted ground truth coordinates to pixel coordinates. Lastly, you have created a Folium map showing ground truth points, pixel footprints and two radar backscatter layers.
12. Resources and references¶
Authors: Julia White, Alex Lewandowski
- Bonatti, E., Gasperini, E., Vigliotti, L., Lupi, L., Vaselli, O., Polonia, A., & Gasperini, L. (2017). Lake Afrera, a structural depression in the Northern Afar Rift (Red Sea). Heliyon, 3(5), e00301. 10.1016/j.heliyon.2017.e00301