asf_search is an open source Python package for searching SAR data archived at ASF. This notebook demonstrates how to search NISAR GCOV with asf_search, stream them directly over HTTPS or from S3 storage with s3fs, and load them into xarray data structures.
NISAR data products can be very large. It may be helpful to access the data directly from their S3 storage. This allows you to lazily load data into xarray data structures, subset, and perform operations on them with xarray, and then only save the data you need to memory. This avoids downloading many large products to a storage volume, only to subset and delete most of them. The caveat is that you must have enough RAM to hold your final subset and also run your workflow.
Overview¶
1. Prerequisites¶
| Prerequisite | Importance | Notes |
|---|---|---|
| The software environment for this cookbook must be installed | Necessary |
Rough Notebook Time Estimate: 10 minutes
2. Search for data¶
Use asf_search to find GCOV data.
2a. Perform an asf_search.search() to identify your desired product URLs¶
import os
import asf_search as asf
from datetime import datetime
from getpass import getpass
import warnings
warnings.filterwarnings(
"ignore",
message="Parsing dates involving a day of month without a year specified",
)
session = asf.ASFSession()
start_date = datetime(2025, 11, 22)
end_date = datetime(2026, 3, 5)
area_of_interest = "POLYGON((40.9131 12.3904,41.8891 12.3904,41.8891 13.2454,40.9131 13.2454,40.9131 12.3904))" # POINT or POLYGON as WKT (well-known-text)
pattern = r'^(?!.*QA_STATS).*'
opts=asf.ASFSearchOptions(**{
"maxResults": 250,
"intersectsWith": area_of_interest,
"flightDirection": "ASCENDING",
"start": start_date,
"end": end_date,
"processingLevel": [
"GCOV"
],
"dataset": [
"NISAR"
],
"productionConfiguration": [
"PR"
],
'session': session,
})
response = asf.search(opts=opts)
hdf5_urls = response.find_urls(extension='.h5', pattern=pattern, directAccess=False)
print(f"Found {len(hdf5_urls)} GCOV products:")
hdf5_urlsFound 7 GCOV products:
['https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05007_N_F_J_001/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05007_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05009_N_F_J_001/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05007_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05007_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_007_172_A_008_2005_DHDH_A_20251216T024619_20251216T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_007_172_A_008_2005_DHDH_A_20251216T024619_20251216T024653_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_008_172_A_008_2005_DHDH_A_20251228T024619_20251228T024654_X05009_N_F_J_001/NISAR_L2_PR_GCOV_008_172_A_008_2005_DHDH_A_20251228T024619_20251228T024654_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_009_172_A_008_2005_DHDH_A_20260109T024620_20260109T024654_X05009_N_F_J_001/NISAR_L2_PR_GCOV_009_172_A_008_2005_DHDH_A_20260109T024620_20260109T024654_X05009_N_F_J_001.h5']2b. Retain only the URL for the most recent version of each product in the search results¶
Data is occasionally re-released with an updated version. Versions are recorded as a Composite Release Identifier (CRID) in a product’s filename. We can use the CRID to retain only the most recent version of each product in the list of URLs.
import re
pattern = re.compile(r"(NISAR_L2_PR_GCOV(?:_[^_]+){9})_(X\d{5})")
latest_version_dict = {}
for url in hdf5_urls:
m = pattern.search(url)
if not m:
continue
product, crid = m.groups()
if product not in latest_version_dict or crid > latest_version_dict[product][0]:
latest_version_dict[product] = (crid, url)
hdf5_urls = [i[1] for i in latest_version_dict.values()]
print(f"Retained {len(hdf5_urls)} GCOV products:")
hdf5_urlsRetained 5 GCOV products:
['https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05009_N_F_J_001/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_007_172_A_008_2005_DHDH_A_20251216T024619_20251216T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_007_172_A_008_2005_DHDH_A_20251216T024619_20251216T024653_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_008_172_A_008_2005_DHDH_A_20251228T024619_20251228T024654_X05009_N_F_J_001/NISAR_L2_PR_GCOV_008_172_A_008_2005_DHDH_A_20251228T024619_20251228T024654_X05009_N_F_J_001.h5',
'https://nisar.asf.earthdatacloud.nasa.gov/NISAR/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_009_172_A_008_2005_DHDH_A_20260109T024620_20260109T024654_X05009_N_F_J_001/NISAR_L2_PR_GCOV_009_172_A_008_2005_DHDH_A_20260109T024620_20260109T024654_X05009_N_F_J_001.h5']2c. Provide your Earthdata Login (EDL) Bearer Token¶
Both HTTPS and S3 access require an EDL Bearer Token
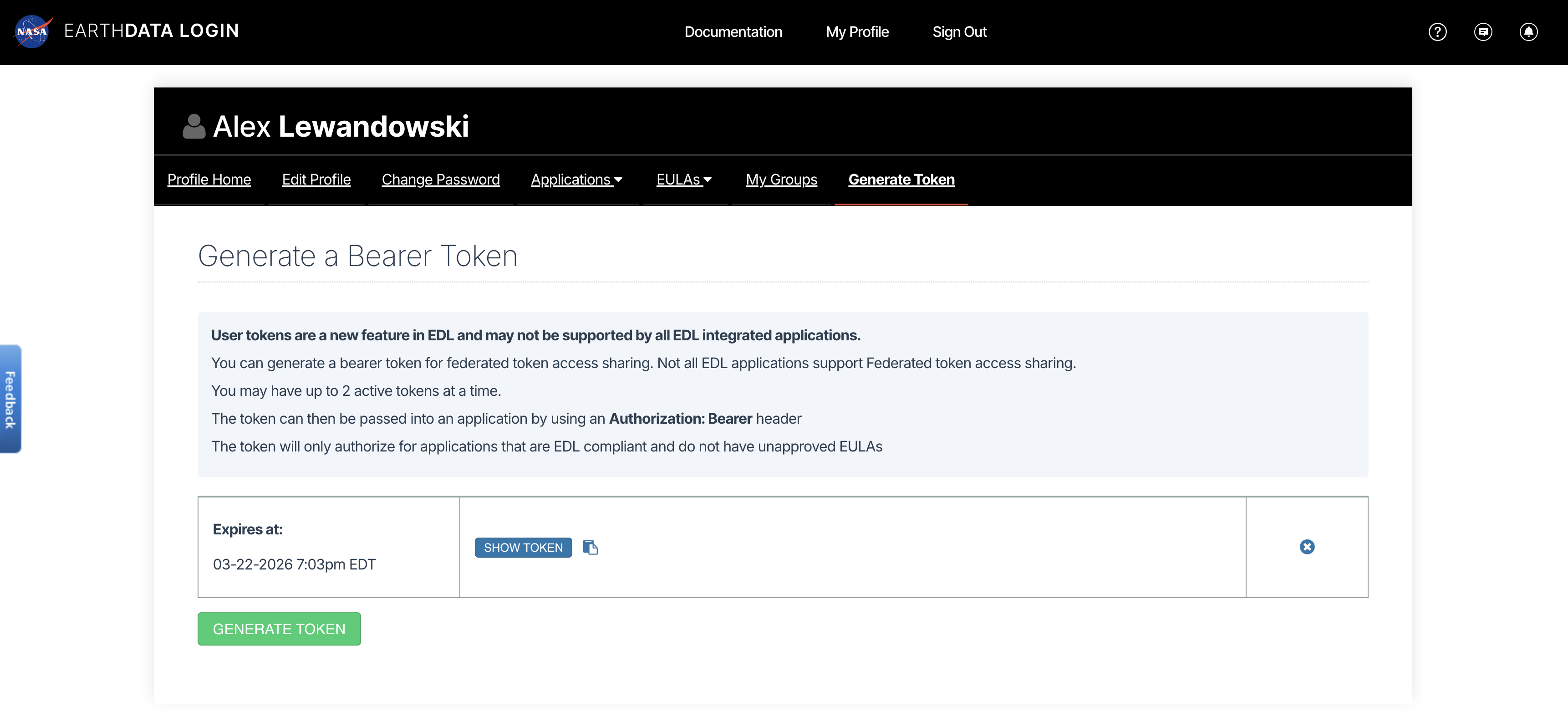
View or generate a Bearer Token in “Generate Token” tab of the Profile page in your Earthdata Login account: https://
from getpass import getpass
token = getpass("Enter your EDL Bearer Token")Enter your EDL Bearer Token ········
3. Load a single GCOV product with HTTPS¶
This example loads the data in a Python with statement.
3a. Open a GCOV file in a with block and compute the mean of an HHHH spatial subset¶
Note that any computation on the data (in this case a subset mean) must be performed inside the with block.
%%time
import asf_search as asf
import fsspec
import rioxarray
import xarray as xr
fs = fsspec.filesystem(
"http",
headers = {"Authorization": f"Bearer {token}"},
block_size = 16 * 512 * 512,
)
with fs.open(hdf5_urls[0], "rb") as f:
dt = xr.open_datatree(
f,
engine="h5netcdf",
decode_timedelta=False,
phony_dims="access",
chunks="auto",
)
### Perform any calculations and save any computed results for future access here, inside the `with` block ###
frequencyA = dt["/science/LSAR/GCOV/grids/frequencyA"] # access the frequency A data
projection = frequencyA.projection.attrs['epsg_code'].item() # access the GCOV product's projection
hhhh = frequencyA.HHHH # access frequency A's HHHH band
hhhh = hhhh.rio.write_crs(projection) # write the project to the HHHH data for easy lat/lon subsetting
# subset the data
subset_hhhh = hhhh.rio.clip_box(
minx=40.8463, miny=13.2553,
maxx=40.8574, maxy=13.2684,
crs="EPSG:4326"
)
# save the mean of HHHH subset for use outside of the `with` block
subset_hhhh_mean = subset_hhhh.mean().to_numpy().item()
print(f'subset_hhhh_mean: {subset_hhhh_mean}\n')subset_hhhh_mean: 0.07500407099723816
CPU times: user 2.6 s, sys: 509 ms, total: 3.11 s
Wall time: 41.8 s
4. Load a single GCOV product from the S3 bucket¶
This example loads the data in a Python with statement.
4a. Convert the hdf5 URLs into S3 urls¶
from urllib.parse import urlparse
bucket = "sds-n-cumulus-prod-nisar-products"
s3_urls = [f"s3://{bucket}/{'/'.join(urlparse(url).path.split('/')[2:])}" for url in hdf5_urls]
s3_urls['s3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05009_N_F_J_001/NISAR_L2_PR_GCOV_005_172_A_008_2005_DHDH_A_20251122T024618_20251122T024652_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_006_172_A_008_2005_DHDH_A_20251204T024618_20251204T024653_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_007_172_A_008_2005_DHDH_A_20251216T024619_20251216T024653_X05009_N_F_J_001/NISAR_L2_PR_GCOV_007_172_A_008_2005_DHDH_A_20251216T024619_20251216T024653_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_008_172_A_008_2005_DHDH_A_20251228T024619_20251228T024654_X05009_N_F_J_001/NISAR_L2_PR_GCOV_008_172_A_008_2005_DHDH_A_20251228T024619_20251228T024654_X05009_N_F_J_001.h5',
's3://sds-n-cumulus-prod-nisar-products/NISAR_L2_GCOV_BETA_V1/NISAR_L2_PR_GCOV_009_172_A_008_2005_DHDH_A_20260109T024620_20260109T024654_X05009_N_F_J_001/NISAR_L2_PR_GCOV_009_172_A_008_2005_DHDH_A_20260109T024620_20260109T024654_X05009_N_F_J_001.h5']4b. Use your EDL Bearer Token to get S3 bucket access credentials¶
The S3 credentials acquired below expire after 1 hour.
import json
import s3fs
import urllib
prefix = "NISAR_L2_GCOV_BETA_V1"
event = {
"CredentialsEndpoint": "https://nisar.asf.earthdatacloud.nasa.gov/s3credentials",
"BearerToken": token,
"Bucket": bucket,
"Prefix": prefix,
}
# Get temporary download credentials
tea_url = event["CredentialsEndpoint"]
bearer_token = event["BearerToken"]
req = urllib.request.Request(
url=tea_url,
headers={"Authorization": f"Bearer {bearer_token}"}
)
with urllib.request.urlopen(req) as f:
creds = json.loads(f.read().decode())4c. Open a GCOV file in a with block and compute the mean of an HHHH spatial subset¶
Notes on Chunking¶
This notebook utilizes Dask chunking to break large arrays into smaller blocks. Chunking allows users to inspect and process subsets of a dataset without loading entire arrays into memory at once. This notebook uses a chunk size of 8 MB.
For additional benchmarking and recommendations for NISAR direct-access workflows, see Henry Rodman’s Reading NISAR granules directly from S3.
%%time
import xarray as xr
import s3fs
import rioxarray
s3_url = s3_urls[0]
fs = s3fs.S3FileSystem(
key=creds["accessKeyId"],
secret=creds["secretAccessKey"],
token=creds["sessionToken"],
)
kwargs = {
"cache_type": "background",
"block_size": 8 * 512 * 512, # 8 MB
}
with fs.open(s3_url, "rb", **kwargs) as f:
dt = xr.open_datatree(
f,
engine="h5netcdf",
decode_timedelta=False,
chunks="auto",
phony_dims="access"
)
### Perform any calculations and save any computed results for future access here, inside the `with` block ###
frequencyA = dt["/science/LSAR/GCOV/grids/frequencyA"] # access the frequency A data
projection = frequencyA.projection.attrs['epsg_code'].item() # access the GCOV product's projection
hhhh = frequencyA.HHHH # access frequency A's HHHH band
hhhh = hhhh.rio.write_crs(projection) # write the project to the HHHH data for easy lat/lon subsetting
# subset the data
subset_hhhh = hhhh.rio.clip_box(
minx=40.8463, miny=13.2553,
maxx=40.8574, maxy=13.2684,
crs="EPSG:4326"
)
# save the mean of HHHH subset for use outside of the `with` block
subset_hhhh_mean = subset_hhhh.mean().to_numpy().item()
print(f'subset_hhhh_mean: {subset_hhhh_mean}\n')subset_hhhh_mean: 0.07500407099723816
CPU times: user 991 ms, sys: 70.4 ms, total: 1.06 s
Wall time: 2.08 s
4d. View the DataTree¶
You can look at the DataTree outside the with block, but it only contains pointers to the data and the file is closed, so you can no longer access any of the data values to which it points
dt4e. Try (and fail) to compute a value in the DataTree¶
The file was closed upon exiting the with block so trying to compute a value from the DataTree will fail with a ValueError: I/O operation on closed file.
dt.compute()---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
Cell In[9], line 1
----> 1 dt.compute()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/datatree.py:2458, in DataTree.compute(self, **kwargs)
2434 """Manually trigger loading and/or computation of this datatree's data
2435 from disk or a remote source into memory and return a new datatree.
2436 Unlike load, the original datatree is left unaltered.
(...) 2455 dask.compute
2456 """
2457 new = self.copy(deep=False)
-> 2458 return new.load(**kwargs)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/datatree.py:2416, in DataTree.load(self, **kwargs)
2413 chunkmanager = get_chunked_array_type(*flat_lazy_data.values())
2415 # evaluate all the chunked arrays simultaneously
-> 2416 evaluated_data: tuple[np.ndarray[Any, Any], ...] = chunkmanager.compute(
2417 *flat_lazy_data.values(), **kwargs
2418 )
2420 for (path, var_name), data in zip(
2421 flat_lazy_data, evaluated_data, strict=False
2422 ):
2423 self[path].variables[var_name].data = data
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/namedarray/daskmanager.py:85, in DaskManager.compute(self, *data, **kwargs)
80 def compute(
81 self, *data: Any, **kwargs: Any
82 ) -> tuple[np.ndarray[Any, _DType_co], ...]:
83 from dask.array import compute
---> 85 return compute(*data, **kwargs)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/dask/base.py:686, in compute(traverse, optimize_graph, scheduler, get, *args, **kwargs)
683 expr = expr.optimize()
684 keys = list(flatten(expr.__dask_keys__()))
--> 686 results = schedule(expr, keys, **kwargs)
688 return repack(results)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:686, in ImplicitToExplicitIndexingAdapter.__array__(self, dtype, copy)
682 def __array__(
683 self, dtype: DTypeLike | None = None, /, *, copy: bool | None = None
684 ) -> np.ndarray:
685 if Version(np.__version__) >= Version("2.0.0"):
--> 686 return np.asarray(self.get_duck_array(), dtype=dtype, copy=copy)
687 else:
688 return np.asarray(self.get_duck_array(), dtype=dtype)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:691, in ImplicitToExplicitIndexingAdapter.get_duck_array(self)
690 def get_duck_array(self):
--> 691 return self.array.get_duck_array()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:924, in CopyOnWriteArray.get_duck_array(self)
923 def get_duck_array(self):
--> 924 return self.array.get_duck_array()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:764, in LazilyIndexedArray.get_duck_array(self)
761 from xarray.backends.common import BackendArray
763 if isinstance(self.array, BackendArray):
--> 764 array = self.array[self.key]
765 else:
766 array = apply_indexer(self.array, self.key)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/backends/h5netcdf_.py:67, in H5NetCDFArrayWrapper.__getitem__(self, key)
66 def __getitem__(self, key):
---> 67 return indexing.explicit_indexing_adapter(
68 key, self.shape, indexing.IndexingSupport.OUTER_1VECTOR, self._getitem
69 )
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:1156, in explicit_indexing_adapter(key, shape, indexing_support, raw_indexing_method)
1134 """Support explicit indexing by delegating to a raw indexing method.
1135
1136 Outer and/or vectorized indexers are supported by indexing a second time
(...) 1153 Indexing result, in the form of a duck numpy-array.
1154 """
1155 raw_key, numpy_indices = decompose_indexer(key, shape, indexing_support)
-> 1156 result = raw_indexing_method(raw_key.tuple)
1157 if numpy_indices.tuple:
1158 # index the loaded duck array
1159 indexable = as_indexable(result)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/backends/h5netcdf_.py:74, in H5NetCDFArrayWrapper._getitem(self, key)
72 with self.datastore.lock:
73 array = self.get_array(needs_lock=False)
---> 74 return array[key]
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/h5netcdf/core.py:729, in BaseVariable.__getitem__(self, key)
727 return h5ds[key].view(view)
728 else:
--> 729 return h5ds[key]
File h5py/_objects.pyx:54, in h5py._objects.with_phil.wrapper()
File h5py/_objects.pyx:55, in h5py._objects.with_phil.wrapper()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/h5py/_hl/dataset.py:840, in Dataset.__getitem__(self, args, new_dtype)
838 if self._fast_read_ok and (new_dtype is None):
839 try:
--> 840 return self._fast_reader.read(args)
841 except TypeError:
842 pass # Fall back to Python read pathway below
File h5py/_selector.pyx:375, in h5py._selector.Reader.read()
File h5py/h5fd.pyx:159, in h5py.h5fd.H5FD_fileobj_read()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/fsspec/spec.py:2140, in AbstractBufferedFile.readinto(self, b)
2135 """mirrors builtin file's readinto method
2136
2137 https://docs.python.org/3/library/io.html#io.RawIOBase.readinto
2138 """
2139 out = memoryview(b).cast("B")
-> 2140 data = self.read(out.nbytes)
2141 out[: len(data)] = data
2142 return len(data)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/fsspec/spec.py:2118, in AbstractBufferedFile.read(self, length)
2116 length = self.size - self.loc
2117 if self.closed:
-> 2118 raise ValueError("I/O operation on closed file.")
2119 if length == 0:
2120 # don't even bother calling fetch
2121 return b""
ValueError: I/O operation on closed file.However, subset_hhhh_mean was computed and saved to memory inside the with block, so we can still see its value¶
subset_hhhh_mean5. Load multiple GCOV Products at once¶
5a. Iterate through a list of HDF5 S3 bucket URLs, and open the /science/LSAR/GCOV/grids/frequencyA for each¶
This leaves the files open for later use, which means you should manually close them when finished in order to prevent memory leaks.
%%time
import xarray as xr
import rioxarray
# Explore the DataTree rendering above in Step 4 for a complete list of available groups
group_path = "/science/LSAR/GCOV/grids/frequencyA" # change this to any GCOV HDF5 group you wish
kwargs = {
"cache_type": "background",
"block_size": 16 * 1024 * 1024, # 16 MB
}
files = [fs.open(url, "rb", **kwargs) for url in s3_urls]
datatrees = [
xr.open_datatree(
f,
engine="h5netcdf",
decode_timedelta=False,
phony_dims="access",
chunks="auto",
group=group_path,
)
for f in files
]CPU times: user 836 ms, sys: 413 ms, total: 1.25 s
Wall time: 5.52 s
5b. Since the files remain open, we can access still access them and perform computations on their contents¶
Iterate through the DataTrees, calculating subset HHHH mean values
for tree in datatrees:
projection = tree.projection.attrs['epsg_code'].item()
hhhh = tree.HHHH
hhhh = hhhh.rio.write_crs(projection)
subset_hhhh = hhhh.rio.clip_box(
minx=40.8463, miny=13.2553,
maxx=40.8574, maxy=13.2684,
crs="EPSG:4326"
)
subset_hhhh_mean = subset_hhhh.mean().to_numpy().item()
print(subset_hhhh_mean)0.07500407099723816
0.07547999173402786
0.0776161402463913
0.07449550926685333
0.07419200986623764
6. Stream a time series¶
Use the multiple files we opened in Step 5 to create a GCOV time series
6a. Define a function to extract datetimes for a time dimension¶
import re
from urllib.parse import urlparse
from datetime import datetime
from pathlib import PurePosixPath
NISAR_TS_RE = re.compile(r"_(\d{8}T\d{6})_")
def nisar_start_time_from_url(s3_url: str) -> datetime:
path = urlparse(s3_url).path
name = PurePosixPath(path).name
m = NISAR_TS_RE.search(name)
if not m:
raise ValueError(f"No NISAR timestamp found in: {s3_url}")
return datetime.strptime(m.group(1), "%Y%m%dT%H%M%S")6b. Create a list of datetimes for a time dimension¶
dts = [nisar_start_time_from_url(url) for url in s3_urls]
dts[datetime.datetime(2025, 11, 22, 2, 46, 18),
datetime.datetime(2025, 12, 4, 2, 46, 18),
datetime.datetime(2025, 12, 16, 2, 46, 19),
datetime.datetime(2025, 12, 28, 2, 46, 19),
datetime.datetime(2026, 1, 9, 2, 46, 20)]6c. Create an xarray.DataArray containing a time dimension for each open GCOV group¶
dataarrays = [
tree.ds.assign_coords(time=dt).expand_dims(time=1)
for dt, tree in zip(dts, datatrees)
]
dataarrays[0].timefor da in dataarrays:
print(da.dims)FrozenMappingWarningOnValuesAccess({'time': 1, 'yCoordinates': 16704, 'xCoordinates': 17064, 'phony_dim_0': 2})
FrozenMappingWarningOnValuesAccess({'time': 1, 'yCoordinates': 16704, 'xCoordinates': 17064, 'phony_dim_0': 2})
FrozenMappingWarningOnValuesAccess({'time': 1, 'yCoordinates': 16704, 'xCoordinates': 17064, 'phony_dim_0': 2})
FrozenMappingWarningOnValuesAccess({'time': 1, 'yCoordinates': 16704, 'xCoordinates': 17064, 'phony_dim_0': 2})
FrozenMappingWarningOnValuesAccess({'time': 1, 'yCoordinates': 16704, 'xCoordinates': 17064, 'phony_dim_0': 2})
6d. Concatenate the xarray.DataArrays into a single xarray.Dataset time series¶
ts = xr.concat(dataarrays, dim="time")
tsfor f in files:
f.close()7b. With closed files, we can no longer compute values in the xarray data structures that we have created¶
The cell below will return a ValueError: I/O operation on closed file.
ts.compute()---------------------------------------------------------------------------
ValueError Traceback (most recent call last)
Cell In[18], line 1
----> 1 ts.compute()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/dataset.py:798, in Dataset.compute(self, **kwargs)
768 """Trigger loading data into memory and return a new dataset.
769
770 Data will be computed and/or loaded from disk or a remote source.
(...) 795 Variable.compute
796 """
797 new = self.copy(deep=False)
--> 798 return new.load(**kwargs)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/dataset.py:564, in Dataset.load(self, **kwargs)
561 chunkmanager = get_chunked_array_type(*chunked_data.values())
563 # evaluate all the chunked arrays simultaneously
--> 564 evaluated_data: tuple[np.ndarray[Any, Any], ...] = chunkmanager.compute(
565 *chunked_data.values(), **kwargs
566 )
568 for k, data in zip(chunked_data, evaluated_data, strict=False):
569 self.variables[k].data = data
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/namedarray/daskmanager.py:85, in DaskManager.compute(self, *data, **kwargs)
80 def compute(
81 self, *data: Any, **kwargs: Any
82 ) -> tuple[np.ndarray[Any, _DType_co], ...]:
83 from dask.array import compute
---> 85 return compute(*data, **kwargs)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/dask/base.py:686, in compute(traverse, optimize_graph, scheduler, get, *args, **kwargs)
683 expr = expr.optimize()
684 keys = list(flatten(expr.__dask_keys__()))
--> 686 results = schedule(expr, keys, **kwargs)
688 return repack(results)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:686, in ImplicitToExplicitIndexingAdapter.__array__(self, dtype, copy)
682 def __array__(
683 self, dtype: DTypeLike | None = None, /, *, copy: bool | None = None
684 ) -> np.ndarray:
685 if Version(np.__version__) >= Version("2.0.0"):
--> 686 return np.asarray(self.get_duck_array(), dtype=dtype, copy=copy)
687 else:
688 return np.asarray(self.get_duck_array(), dtype=dtype)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:691, in ImplicitToExplicitIndexingAdapter.get_duck_array(self)
690 def get_duck_array(self):
--> 691 return self.array.get_duck_array()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:924, in CopyOnWriteArray.get_duck_array(self)
923 def get_duck_array(self):
--> 924 return self.array.get_duck_array()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:764, in LazilyIndexedArray.get_duck_array(self)
761 from xarray.backends.common import BackendArray
763 if isinstance(self.array, BackendArray):
--> 764 array = self.array[self.key]
765 else:
766 array = apply_indexer(self.array, self.key)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/backends/h5netcdf_.py:67, in H5NetCDFArrayWrapper.__getitem__(self, key)
66 def __getitem__(self, key):
---> 67 return indexing.explicit_indexing_adapter(
68 key, self.shape, indexing.IndexingSupport.OUTER_1VECTOR, self._getitem
69 )
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/core/indexing.py:1156, in explicit_indexing_adapter(key, shape, indexing_support, raw_indexing_method)
1134 """Support explicit indexing by delegating to a raw indexing method.
1135
1136 Outer and/or vectorized indexers are supported by indexing a second time
(...) 1153 Indexing result, in the form of a duck numpy-array.
1154 """
1155 raw_key, numpy_indices = decompose_indexer(key, shape, indexing_support)
-> 1156 result = raw_indexing_method(raw_key.tuple)
1157 if numpy_indices.tuple:
1158 # index the loaded duck array
1159 indexable = as_indexable(result)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/xarray/backends/h5netcdf_.py:74, in H5NetCDFArrayWrapper._getitem(self, key)
72 with self.datastore.lock:
73 array = self.get_array(needs_lock=False)
---> 74 return array[key]
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/h5netcdf/core.py:729, in BaseVariable.__getitem__(self, key)
727 return h5ds[key].view(view)
728 else:
--> 729 return h5ds[key]
File h5py/_objects.pyx:54, in h5py._objects.with_phil.wrapper()
File h5py/_objects.pyx:55, in h5py._objects.with_phil.wrapper()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/h5py/_hl/dataset.py:840, in Dataset.__getitem__(self, args, new_dtype)
838 if self._fast_read_ok and (new_dtype is None):
839 try:
--> 840 return self._fast_reader.read(args)
841 except TypeError:
842 pass # Fall back to Python read pathway below
File h5py/_selector.pyx:375, in h5py._selector.Reader.read()
File h5py/h5fd.pyx:159, in h5py.h5fd.H5FD_fileobj_read()
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/fsspec/spec.py:2140, in AbstractBufferedFile.readinto(self, b)
2135 """mirrors builtin file's readinto method
2136
2137 https://docs.python.org/3/library/io.html#io.RawIOBase.readinto
2138 """
2139 out = memoryview(b).cast("B")
-> 2140 data = self.read(out.nbytes)
2141 out[: len(data)] = data
2142 return len(data)
File ~/NISAR_GCOV_Cookbook/.pixi/envs/isce3/lib/python3.13/site-packages/fsspec/spec.py:2118, in AbstractBufferedFile.read(self, length)
2116 length = self.size - self.loc
2117 if self.closed:
-> 2118 raise ValueError("I/O operation on closed file.")
2119 if length == 0:
2120 # don't even bother calling fetch
2121 return b""
ValueError: I/O operation on closed file.8. Summary¶
You now have the tools and knowledge that you need to search with asf_search, generate temporary S3 bucket credentials from an Earthdata Login Bearer Token, stream data from S3 with s3fs, and load them into xarray data structures.